3D Example: GDDEGP on Hartmann3 Function with Multiple Directional Derivatives#
This example demonstrates GDDEGP on a 3-variable function using gradient-aligned and perpendicular directional derivatives at each training point.
Configuration#
import numpy as np
import pyoti.sparse as oti
import itertools
from jetgp.full_gddegp.gddegp import gddegp
from scipy.stats import qmc
import jetgp.utils as utils
n_order = 1
n_bases = 3
num_directions_per_point = 3 # gradient + 2 perpendicular directions
num_training_pts = 50
domain_bounds = ((0.0, 1.0), (0.0, 1.0), (0.0, 1.0))
normalize_data = True
kernel = "SE"
kernel_type = "anisotropic"
n_restarts = 15
swarm_size = 200
random_seed = 42
np.random.seed(random_seed)
print("GDDEGP 3D Example: Hartmann3 Function")
print(f"Training points: {num_training_pts}")
print(f"Directions per point: {num_directions_per_point}")
print(f"Total observations: {num_training_pts * (1 + num_directions_per_point)}")
GDDEGP 3D Example: Hartmann3 Function
Training points: 50
Directions per point: 3
Total observations: 200
Define Hartmann3 Function#
def hartmann3(X, alg=np):
"""3D Hartmann function - standard optimization benchmark."""
alpha = np.array([1.0, 1.2, 3.0, 3.2])
A = np.array([[3.0, 10, 30],
[0.1, 10, 35],
[3.0, 10, 30],
[0.1, 10, 35]])
P = 1e-4 * np.array([[3689, 1170, 2673],
[4699, 4387, 7470],
[1091, 8732, 5547],
[381, 5743, 8828]])
result = alg.zeros(X.shape[0]) if hasattr(alg, 'zeros') else np.zeros(X.shape[0])
for i in range(4):
inner = alg.zeros(X.shape[0]) if hasattr(alg, 'zeros') else np.zeros(X.shape[0])
for j in range(3):
inner = inner + A[i, j] * (X[:, j] - P[i, j])**2
result = result - alpha[i] * alg.exp(-inner)
return result
def hartmann3_gradient(x1, x2, x3):
"""Analytical gradient of Hartmann3 function."""
alpha = np.array([1.0, 1.2, 3.0, 3.2])
A = np.array([[3.0, 10, 30],
[0.1, 10, 35],
[3.0, 10, 30],
[0.1, 10, 35]])
P = 1e-4 * np.array([[3689, 1170, 2673],
[4699, 4387, 7470],
[1091, 8732, 5547],
[381, 5743, 8828]])
grad = np.zeros(3)
X = np.array([x1, x2, x3])
for i in range(4):
inner = np.sum(A[i, :] * (X - P[i, :])**2)
exp_term = np.exp(-inner)
for j in range(3):
grad[j] += 2 * alpha[i] * A[i, j] * (X[j] - P[i, j]) * exp_term
return grad
Gram-Schmidt Orthonormalization#
def gram_schmidt_orthonormal(v1):
"""
Generate 2 orthonormal vectors perpendicular to v1.
v1 should be a unit vector.
"""
# Find a vector not parallel to v1
if abs(v1[0]) < 0.9:
v2_init = np.array([1.0, 0.0, 0.0])
else:
v2_init = np.array([0.0, 1.0, 0.0])
# First perpendicular vector
v2 = v2_init - np.dot(v2_init, v1) * v1
v2 = v2 / np.linalg.norm(v2)
# Second perpendicular vector
v3 = np.cross(v1, v2)
v3 = v3 / np.linalg.norm(v3)
return v2, v3
Generate Training Data#
def generate_training_data():
"""Generate GDDEGP training data with multiple rays per point."""
# Latin Hypercube Sampling
sampler = qmc.LatinHypercube(d=n_bases, seed=random_seed)
unit_samples = sampler.random(n=num_training_pts)
X_train = qmc.scale(unit_samples,
[b[0] for b in domain_bounds],
[b[1] for b in domain_bounds])
print("Generating directional derivatives...")
# Create rays: gradient + 2 perpendicular directions
rays_list = [[] for _ in range(num_directions_per_point)]
for idx, (x1, x2, x3) in enumerate(X_train):
# Compute gradient direction
grad = hartmann3_gradient(x1, x2, x3)
grad_norm = np.linalg.norm(grad)
if grad_norm > 1e-10:
ray_grad = grad / grad_norm
else:
ray_grad = np.array([1.0, 0.0, 0.0])
# Generate perpendicular directions
ray_perp1, ray_perp2 = gram_schmidt_orthonormal(ray_grad)
rays_list[0].append(ray_grad.reshape(-1, 1))
rays_list[1].append(ray_perp1.reshape(-1, 1))
rays_list[2].append(ray_perp2.reshape(-1, 1))
if idx < 3: # Show first few
print(f" Point {idx}: grad={ray_grad}")
# Apply global perturbations
X_pert = oti.array(X_train)
for i in range(num_directions_per_point):
e_tag = oti.e(i+1, order=n_order)
for j in range(len(rays_list[i])):
perturbation = oti.array(rays_list[i][j]) * e_tag
X_pert[j, :] += perturbation.T
# Evaluate with hypercomplex AD
f_hc = hartmann3(X_pert, alg=oti)
# Truncate cross-derivatives
for combo in itertools.combinations(range(1, num_directions_per_point+1), 2):
f_hc = f_hc.truncate(combo)
# Extract derivatives
y_train_list = [f_hc.real.reshape(-1, 1)]
der_indices_to_extract = [ [ [[i, 1]] for i in range(1, num_directions_per_point+1)]]
for group in der_indices_to_extract:
for sub_group in group:
y_train_list.append(f_hc.get_deriv(sub_group).reshape(-1, 1))
print(f"Data structure:")
print(f" Function values: {y_train_list[0].shape}")
for i in range(1, len(y_train_list)):
print(f" Direction {i} derivatives: {y_train_list[i].shape}")
return {'X_train': X_train,
'y_train_list': y_train_list,
'rays_list': rays_list,
'der_indices': der_indices_to_extract}
training_data = generate_training_data()
Generating directional derivatives...
Point 0: grad=[ 0.13366276 -0.03709979 -0.9903322 ]
Point 1: grad=[0.28965432 0.082479 0.95357097]
Point 2: grad=[0.01400605 0.23698152 0.97141319]
Data structure:
Function values: (50, 1)
Direction 1 derivatives: (50, 1)
Direction 2 derivatives: (50, 1)
Direction 3 derivatives: (50, 1)
Train GDDEGP Model#
def train_model(training_data):
"""Initialize and train GDDEGP model."""
print("Training GDDEGP model...")
rays_array = [np.hstack(training_data['rays_list'][i])
for i in range(num_directions_per_point)]
derivative_locations = []
for i in range(len(training_data['der_indices'])):
for j in range(len(training_data['der_indices'][i])):
derivative_locations.append([i for i in range(len(training_data['X_train']))])
gp_model = gddegp(
training_data['X_train'],
training_data['y_train_list'],
n_order=1,
rays_list=rays_array,
der_indices=training_data['der_indices'],
derivative_locations = derivative_locations,
normalize=normalize_data,
kernel=kernel,
kernel_type=kernel_type
)
print("Optimizing hyperparameters...")
params = gp_model.optimize_hyperparameters(
optimizer='pso',
pop_size = 100,
n_generations = 30,
local_opt_every = 30,
debug = True
)
return gp_model, params
gp_model, params = train_model(training_data)
Training GDDEGP model...
Optimizing hyperparameters...
Best after iteration 1: [-1.13578663 -0.03183975 0.70908436 0.55759786 -6.62473884] 475.0879328265031
Best after iteration 2: [-1.13578663 -0.03183975 0.70908436 0.55759786 -6.62473884] 475.0879328265031
New best for swarm at iteration 3: [ 0.06008182 0.14651738 0.16020523 0.16618446 -10.1958612 ] 291.02496846212415
Best after iteration 3: [ 0.06008182 0.14651738 0.16020523 0.16618446 -10.1958612 ] 291.02496846212415
New best for swarm at iteration 4: [-0.2815003 -0.26037333 0.44382445 0.47166394 -8.26812236] 286.37708414440465
New best for swarm at iteration 4: [-0.49009976 0.06745064 0.08122518 1.05028914 -6.06492915] 284.4406567925401
Best after iteration 4: [-0.49009976 0.06745064 0.08122518 1.05028914 -6.06492915] 284.4406567925401
New best for swarm at iteration 5: [-0.67419812 -0.02493449 0.45348343 0.62015136 -8.65039042] 279.80001963716524
New best for swarm at iteration 5: [-0.07903037 -0.12040193 0.09115031 0.35779158 -6.11909554] 200.88799679053966
New best for swarm at iteration 5: [-0.49835435 -0.03610454 0.09872779 0.82090678 -8.81026187] 124.37332631663081
New best for swarm at iteration 5: [ -0.47072759 0.16787616 0.20904911 0.16227639 -10.08375482] 94.08479385453231
Best after iteration 5: [ -0.47072759 0.16787616 0.20904911 0.16227639 -10.08375482] 94.08479385453231
New best for swarm at iteration 6: [-0.40236877 -0.10176391 0.20694244 0.37708764 -9.0793952 ] 27.156822517370188
Best after iteration 6: [-0.40236877 -0.10176391 0.20694244 0.37708764 -9.0793952 ] 27.156822517370188
New best for swarm at iteration 7: [-3.43386289e-01 4.04031233e-03 1.63212116e-01 3.12858064e-02
-7.61513832e+00] -19.654967764682198
Best after iteration 7: [-3.43386289e-01 4.04031233e-03 1.63212116e-01 3.12858064e-02
-7.61513832e+00] -19.654967764682198
Best after iteration 8: [-3.43386289e-01 4.04031233e-03 1.63212116e-01 3.12858064e-02
-7.61513832e+00] -19.654967764682198
New best for swarm at iteration 9: [-0.35619011 0.03753074 0.21161892 -0.13991146 -8.17541646] -33.59153324690723
Best after iteration 9: [-0.35619011 0.03753074 0.21161892 -0.13991146 -8.17541646] -33.59153324690723
New best for swarm at iteration 10: [-0.37901092 -0.06942863 0.19534327 0.09943679 -8.60260551] -35.96025857780916
New best for swarm at iteration 10: [-0.3388882 -0.07226647 0.21655321 -0.07406341 -8.45610894] -37.761788391773194
New best for swarm at iteration 10: [-0.35356233 -0.02825659 0.21170166 -0.15980385 -8.45985944] -48.14464679458982
Best after iteration 10: [-0.35356233 -0.02825659 0.21170166 -0.15980385 -8.45985944] -48.14464679458982
New best for swarm at iteration 11: [-0.34854663 -0.03599816 0.20430775 -0.10850802 -8.17410294] -49.281042523053515
Best after iteration 11: [-0.34854663 -0.03599816 0.20430775 -0.10850802 -8.17410294] -49.281042523053515
New best for swarm at iteration 12: [-0.35580315 -0.01835342 0.20924881 -0.13045739 -8.08027696] -49.48333514640959
Best after iteration 12: [-0.35580315 -0.01835342 0.20924881 -0.13045739 -8.08027696] -49.48333514640959
New best for swarm at iteration 13: [-0.35322608 -0.02669736 0.20543192 -0.11740673 -8.05373262] -49.68711853424347
New best for swarm at iteration 13: [-0.34639865 -0.03361641 0.20875908 -0.10489752 -8.27397307] -49.78935084381041
Best after iteration 13: [-0.34639865 -0.03361641 0.20875908 -0.10489752 -8.27397307] -49.78935084381041
New best for swarm at iteration 14: [-0.35764997 -0.03571737 0.21158331 -0.10201048 -8.10560696] -50.034517142754595
Best after iteration 14: [-0.35764997 -0.03571737 0.21158331 -0.10201048 -8.10560696] -50.034517142754595
New best for swarm at iteration 15: [-0.35081987 -0.02830354 0.2129719 -0.13396537 -8.26957478] -50.09713722797679
New best for swarm at iteration 15: [-0.3524496 -0.03170645 0.21321497 -0.1223606 -8.13140763] -50.186086899331286
Best after iteration 15: [-0.3524496 -0.03170645 0.21321497 -0.1223606 -8.13140763] -50.186086899331286
New best for swarm at iteration 16: [-0.35411647 -0.03055337 0.21152798 -0.11807469 -8.11575705] -50.20830976309
Best after iteration 16: [-0.35411647 -0.03055337 0.21152798 -0.11807469 -8.11575705] -50.20830976309
New best for swarm at iteration 17: [-0.35361696 -0.02794712 0.21088713 -0.11546145 -8.13558745] -50.21869884630516
New best for swarm at iteration 17: [-0.35422279 -0.03063639 0.21116141 -0.10891459 -8.08635577] -50.226341737458284
Best after iteration 17: [-0.35422279 -0.03063639 0.21116141 -0.10891459 -8.08635577] -50.226341737458284
Best after iteration 18: [-0.35422279 -0.03063639 0.21116141 -0.10891459 -8.08635577] -50.226341737458284
New best for swarm at iteration 19: [-0.35297493 -0.02983513 0.21377705 -0.12320396 -8.11406661] -50.24501247065544
New best for swarm at iteration 19: [-0.35386444 -0.03083157 0.21266836 -0.11278014 -8.09865097] -50.25153603597673
Best after iteration 19: [-0.35386444 -0.03083157 0.21266836 -0.11278014 -8.09865097] -50.25153603597673
Best after iteration 20: [-0.35386444 -0.03083157 0.21266836 -0.11278014 -8.09865097] -50.25153603597673
New best for swarm at iteration 21: [-0.35360585 -0.03042565 0.2130143 -0.11883835 -8.09690109] -50.251656803956706
Best after iteration 21: [-0.35360585 -0.03042565 0.2130143 -0.11883835 -8.09690109] -50.251656803956706
New best for swarm at iteration 22: [-0.35368562 -0.02991519 0.21258921 -0.11426171 -8.07466459] -50.26063347396982
New best for swarm at iteration 22: [-0.35360328 -0.02872821 0.21294079 -0.11751882 -8.09547114] -50.26580060436751
Best after iteration 22: [-0.35360328 -0.02872821 0.21294079 -0.11751882 -8.09547114] -50.26580060436751
New best for swarm at iteration 23: [-0.35363386 -0.02892215 0.21308072 -0.11721755 -8.09324823] -50.26654821771294
New best for swarm at iteration 23: [-0.353491 -0.02909216 0.21280355 -0.117332 -8.08170678] -50.26687887686964
Best after iteration 23: [-0.353491 -0.02909216 0.21280355 -0.117332 -8.08170678] -50.26687887686964
New best for swarm at iteration 24: [-0.35339968 -0.02922593 0.21358401 -0.11993441 -8.09817317] -50.26748848054635
Best after iteration 24: [-0.35339968 -0.02922593 0.21358401 -0.11993441 -8.09817317] -50.26748848054635
New best for swarm at iteration 25: [-0.35354568 -0.02912478 0.21308101 -0.11729437 -8.08780036] -50.26768420067586
New best for swarm at iteration 25: [-0.35337244 -0.02875332 0.21318135 -0.11934906 -8.09020998] -50.2680950787354
New best for swarm at iteration 25: [-0.35320836 -0.02892774 0.2133559 -0.11841573 -8.08658801] -50.26877301499471
Best after iteration 25: [-0.35320836 -0.02892774 0.2133559 -0.11841573 -8.08658801] -50.26877301499471
New best for swarm at iteration 26: [-0.35315094 -0.02925176 0.21325727 -0.11930526 -8.09369175] -50.26878910691798
New best for swarm at iteration 26: [-0.35335733 -0.02898701 0.2132605 -0.11864715 -8.08689128] -50.269041628296435
New best for swarm at iteration 26: [-0.35307331 -0.0292698 0.21344633 -0.11920884 -8.0877483 ] -50.269314864063716
New best for swarm at iteration 26: [-0.35284149 -0.02888719 0.21345155 -0.11958493 -8.09197102] -50.26940909305765
Best after iteration 26: [-0.35284149 -0.02888719 0.21345155 -0.11958493 -8.09197102] -50.26940909305765
New best for swarm at iteration 27: [-0.35283097 -0.02889607 0.213482 -0.11973084 -8.09234817] -50.26943686266853
New best for swarm at iteration 27: [-0.35317448 -0.02912993 0.21330932 -0.1186572 -8.09008878] -50.26945957652467
Best after iteration 27: [-0.35317448 -0.02912993 0.21330932 -0.1186572 -8.09008878] -50.26945957652467
New best for swarm at iteration 28: [-0.35303421 -0.02892457 0.21340741 -0.11967888 -8.09045859] -50.26950816543396
New best for swarm at iteration 28: [-0.35291719 -0.02903541 0.21337612 -0.11930939 -8.08952154] -50.2695944823885
Best after iteration 28: [-0.35291719 -0.02903541 0.21337612 -0.11930939 -8.08952154] -50.2695944823885
Best after iteration 29: [-0.35291719 -0.02903541 0.21337612 -0.11930939 -8.08952154] -50.2695944823885
New best for swarm at iteration 30: [-0.35298259 -0.0290237 0.21338118 -0.11945538 -8.09133588] -50.269609315513634
Best after iteration 30: [-0.35298259 -0.0290237 0.21338118 -0.11945538 -8.09133588] -50.269609315513634
Stopping: maximum iterations reached --> 30
Evaluate Model#
def evaluate_model(gp_model, params, training_data):
"""Evaluate GDDEGP on test points."""
print("Evaluating model on test set...")
# Generate test points
n_test = 1000
sampler = qmc.LatinHypercube(d=n_bases, seed=random_seed + 1)
unit_test = sampler.random(n=n_test)
X_test = qmc.scale(unit_test,
[b[0] for b in domain_bounds],
[b[1] for b in domain_bounds])
# Predict
y_pred_full = gp_model.predict(
X_test, params,
calc_cov=False, return_deriv=False
)
y_pred = y_pred_full[:n_test]
# Ground truth
y_true = hartmann3(X_test, alg=np)
# Error metrics
nrmse = utils.nrmse(y_true.flatten(), y_pred.flatten())
mae = np.mean(np.abs(y_true - y_pred))
max_error = np.max(np.abs(y_true - y_pred))
print(f"Test set size: {n_test} points")
print(f"Performance metrics:")
print(f" NRMSE: {nrmse:.6f}")
print(f" MAE: {mae:.6f}")
print(f" Max error: {max_error:.6f}")
return nrmse, mae, max_error
nrmse, mae, max_error = evaluate_model(gp_model, params, training_data)
Evaluating model on test set...
Test set size: 1000 points
Performance metrics:
NRMSE: 0.004473
MAE: 0.007262
Max error: 0.157742
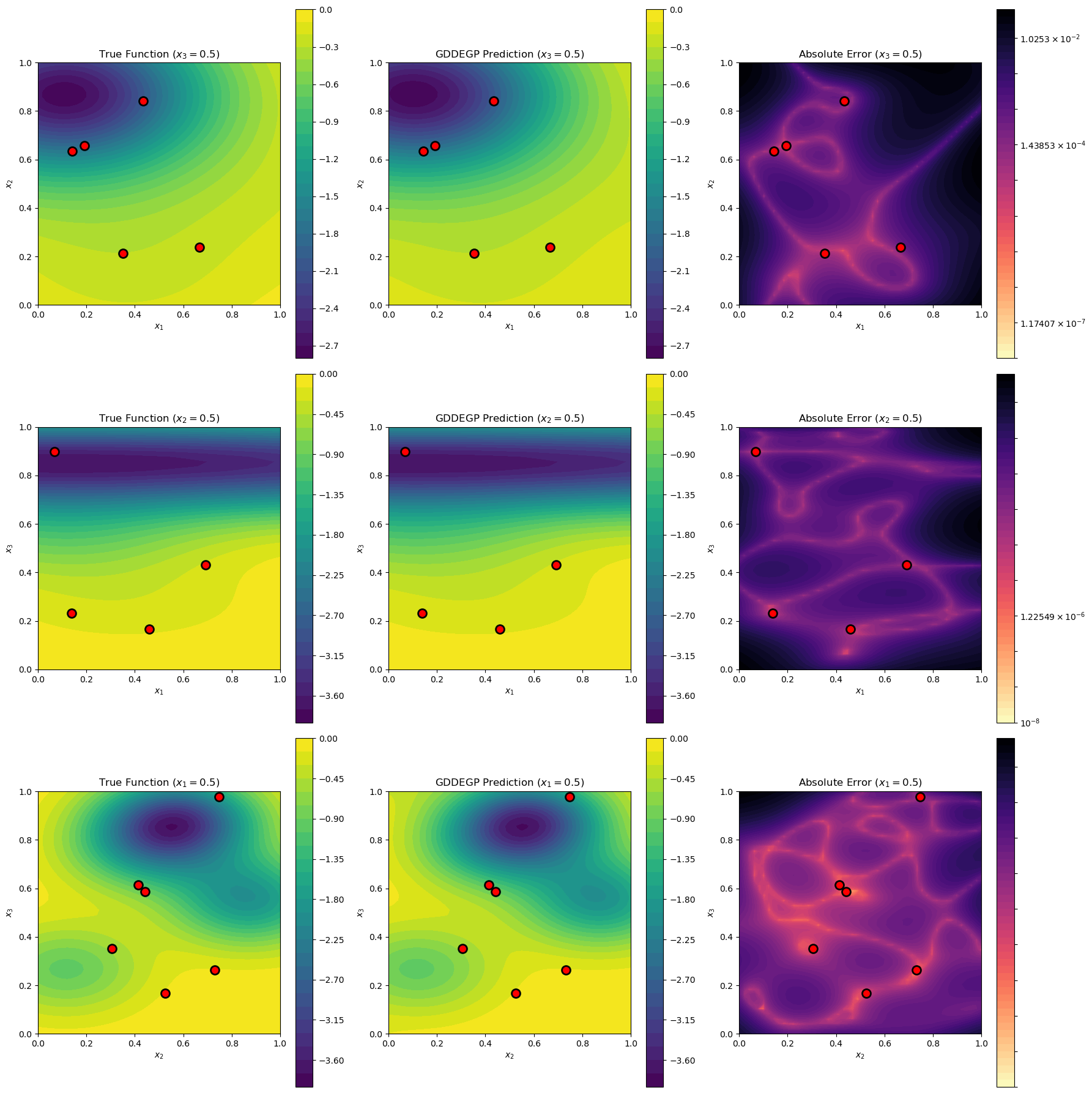
Visualize 2D Slices#
import matplotlib.pyplot as plt
from matplotlib.colors import LogNorm
def visualize_slices(gp_model, params, training_data):
"""Visualize 2D slices through the 3D function."""
print("Generating visualizations...")
X_train = training_data['X_train']
n_points = 50
# Define three slice configurations
slices = [
{'name': '$x_3=0.5$', 'fixed_dim': 2, 'fixed_val': 0.5,
'var_dims': [0, 1], 'labels': ['$x_1$', '$x_2$']},
{'name': '$x_2=0.5$', 'fixed_dim': 1, 'fixed_val': 0.5,
'var_dims': [0, 2], 'labels': ['$x_1$', '$x_3$']},
{'name': '$x_1=0.5$', 'fixed_dim': 0, 'fixed_val': 0.5,
'var_dims': [1, 2], 'labels': ['$x_2$', '$x_3$']}
]
fig, axes = plt.subplots(3, 3, figsize=(18, 18))
for row, slice_info in enumerate(slices):
fixed_dim = slice_info['fixed_dim']
fixed_val = slice_info['fixed_val']
var_dims = slice_info['var_dims']
# Create slice grid
x1_vals = np.linspace(0, 1, n_points)
x2_vals = np.linspace(0, 1, n_points)
X1_grid, X2_grid = np.meshgrid(x1_vals, x2_vals)
# Build full 3D points for this slice
X_slice = np.zeros((n_points * n_points, 3))
X_slice[:, var_dims[0]] = X1_grid.ravel()
X_slice[:, var_dims[1]] = X2_grid.ravel()
X_slice[:, fixed_dim] = fixed_val
# Predict
dummy_ray = np.array([[1.0], [0.0], [0.0]])
rays_pred = [np.hstack([dummy_ray for _ in range(len(X_slice))])
for _ in range(num_directions_per_point)]
y_pred_full = gp_model.predict(X_slice, params,
calc_cov=False, return_deriv=False)
y_pred = y_pred_full[:len(X_slice)].reshape(n_points, n_points)
# Ground truth
y_true = hartmann3(X_slice, alg=np).reshape(n_points, n_points)
# Error
abs_error = np.abs(y_pred - y_true)
abs_error_clipped = np.clip(abs_error, 1e-8, None)
# Filter training points in this slice
tol = 0.05
in_slice = np.abs(X_train[:, fixed_dim] - fixed_val) < tol
X_train_slice = X_train[in_slice]
# Plot 1: True function
ax = axes[row, 0]
c = ax.contourf(X1_grid, X2_grid, y_true, levels=30, cmap='viridis')
if len(X_train_slice) > 0:
ax.scatter(X_train_slice[:, var_dims[0]], X_train_slice[:, var_dims[1]],
c='red', s=100, edgecolors='k', linewidths=2, zorder=5, marker='o')
ax.set_xlabel(slice_info['labels'][0])
ax.set_ylabel(slice_info['labels'][1])
ax.set_title(f"True Function ({slice_info['name']})")
ax.set_aspect('equal')
plt.colorbar(c, ax=ax)
# Plot 2: Prediction
ax = axes[row, 1]
c = ax.contourf(X1_grid, X2_grid, y_pred, levels=30, cmap='viridis')
if len(X_train_slice) > 0:
ax.scatter(X_train_slice[:, var_dims[0]], X_train_slice[:, var_dims[1]],
c='red', s=100, edgecolors='k', linewidths=2, zorder=5, marker='o')
ax.set_xlabel(slice_info['labels'][0])
ax.set_ylabel(slice_info['labels'][1])
ax.set_title(f"GDDEGP Prediction ({slice_info['name']})")
ax.set_aspect('equal')
plt.colorbar(c, ax=ax)
# Plot 3: Error
ax = axes[row, 2]
log_levels = np.logspace(np.log10(abs_error_clipped.min()),
np.log10(abs_error_clipped.max()), num=50)
c = ax.contourf(X1_grid, X2_grid, abs_error_clipped,
levels=log_levels, norm=LogNorm(), cmap='magma_r')
if len(X_train_slice) > 0:
ax.scatter(X_train_slice[:, var_dims[0]], X_train_slice[:, var_dims[1]],
c='red', s=100, edgecolors='k', linewidths=2, zorder=5, marker='o')
ax.set_xlabel(slice_info['labels'][0])
ax.set_ylabel(slice_info['labels'][1])
ax.set_title(f"Absolute Error ({slice_info['name']})")
ax.set_aspect('equal')
plt.colorbar(c, ax=ax)
print(f" Slice {slice_info['name']}: {len(X_train_slice)} training points nearby")
plt.tight_layout()
plt.show()
visualize_slices(gp_model, params, training_data)
Generating visualizations...
Slice $x_3=0.5$: 5 training points nearby
Slice $x_2=0.5$: 4 training points nearby
Slice $x_1=0.5$: 6 training points nearby
Summary#
print("GDDEGP 3D Example Summary")
print("="*70)
print(f"Function: Hartmann3 (3D)")
print(f"Training points: {num_training_pts}")
print(f"Directions per point: {num_directions_per_point} (gradient + 2 perpendicular)")
print(f"Total derivative observations: {num_training_pts * num_directions_per_point}")
print(f"Final NRMSE: {nrmse:.6f}")
print(f"\nKey features demonstrated:")
print(" - 3D function approximation")
print(" - Multiple orthogonal directions per point")
print(" - Gradient-aligned primary direction")
print(" - Gram-Schmidt orthogonalization for perpendicular rays")
print(" - Hypercomplex automatic differentiation")
print(" - 2D slice visualizations through 3D space")
GDDEGP 3D Example Summary
======================================================================
Function: Hartmann3 (3D)
Training points: 50
Directions per point: 3 (gradient + 2 perpendicular)
Total derivative observations: 150
Final NRMSE: 0.004473
Key features demonstrated:
- 3D function approximation
- Multiple orthogonal directions per point
- Gradient-aligned primary direction
- Gram-Schmidt orthogonalization for perpendicular rays
- Hypercomplex automatic differentiation
- 2D slice visualizations through 3D space
